Biotechnology: Principles and Processes
Table of Content |
The use of biology to develop technologies and products for the welfare of human beings is known as Biotechnology. It has various applications in different fields such as Therapeutics, Diagnostics, Processed Food, Waste Management, Energy Production, Genetically Modified Crops etc.
The two core techniques that give rise to modern biotechnology are:
-
Genetic Engineering - Genetic Engineering is defined as the direct manipulation of genome of an organism. It involves the transfer of new genes to improve the function or trait. The most important technique of genetic engineering is gene cloning.
-
Manufacturing of Antibiotics, Vaccines, Drugs etc.
Tools for Genetic Engineering
Restriction Enzymes or Molecular Scissors are used to cut DNA to be inserted into the vector. They used to cut DNA fragments with different ends. There are two types of restriction enzymes - Endonucleases and Exonucleases. Endonucleases cut the DNA in the middle whereas Exonucleases cut at the ends. For Example: ECoR1, Hind III, etc. Restriction Enzymes cut at a specific site on DNA known as Restriction Site.
Ligases are the enzyme that joins the two DNA fragments.
Cloning Vectors
Vector is any DNA molecule that carries a gene of interest to be inserted into the host organism. For Example: Plasmid.
 Plasmid is an autonomously replicating extrachromosomal genetic content present in the bacteria. It is different from the chromosomal DNA. It is used as a vehicle for transfer of gene of interest into the host cell. Plasmid contain origin of replication, site where replication begins when gene of interest enters the host cell. It also contains antibiotic resistance gene.
Plasmid is an autonomously replicating extrachromosomal genetic content present in the bacteria. It is different from the chromosomal DNA. It is used as a vehicle for transfer of gene of interest into the host cell. Plasmid contain origin of replication, site where replication begins when gene of interest enters the host cell. It also contains antibiotic resistance gene.
Following features are required for a cloning vector:
-
Origin of replication
-
Selectable marker to identify transformed cells. Transformed cells are those cells which contain gene of interest. For example, antibiotic resistance gene in plasmid.
-
Cloning sites
Vectors for Cloning in Plants
Agrobacterium tumifaciens, a pathogen of several dicot plants is used as a vector for plants. It  is able to delive a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen. Gene of interest is inserted into T-DNA to transform plant cells with required gene. The Tumor Inducing (Ti) plasmid of Agrobacterium tumifaciens has now been modified into a cloning vector which is no more pathogenic to the plants. Cytokinin and Auxin coding genes in plasmid acts as growth regulator. Opine catabolism gene codes for energy source. Right and left border are needed to transfer T-DNA into the required host plant cell.
is able to delive a piece of DNA known as ‘T-DNA’ to transform normal plant cells into a tumor and direct these tumor cells to produce the chemicals required by the pathogen. Gene of interest is inserted into T-DNA to transform plant cells with required gene. The Tumor Inducing (Ti) plasmid of Agrobacterium tumifaciens has now been modified into a cloning vector which is no more pathogenic to the plants. Cytokinin and Auxin coding genes in plasmid acts as growth regulator. Opine catabolism gene codes for energy source. Right and left border are needed to transfer T-DNA into the required host plant cell.
Process of Recombinant DNA Technology
Steps involved in recombinant DNA technology are as follows:
-
Isolation of the genetic material. There are different methods to isolate DNA content from the bacterial, plant or animal cell. The enzymes used for degradation of wall of the cells are lysozyme (Bacteria), Cellulase (Plant Cell), Chitinase (fungus). RNA is removed by ribonucleases whereas proteins are removed by proteases.
-
Cutting of DNA at specific location. Restriction enzyme or molecular scissors are used to cut DNA at specific locations.
-
PCR of gene of interest. Multiple copies of gene of interest is synthesized in this step.
-
Transfer of gene of interest into specific vector to form a recombinant DNA molecule.
-
Transfer of recombinant DNA molecule into specific host cell.
-
Selection of recombinant cells.
-
Isolation of recombinant cells to obtain recombinant protein.
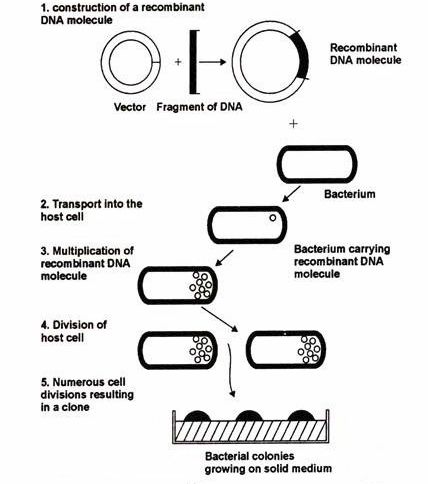
Fig.1. Steps of recombinant DNA technology
Selection of Transformed Cells
There are different methods of selection of transformed cells
After the gene of interest is transferred into host cells, there are two types of cells: Transformed Cells and Non-Transformed cells. Transformed Cells are those cells which contain gene of interest as well as antibiotic resistant gene whereas Non-Transformed Cells do not contain the inserted gene. For this, vector should contain the selectable marker such as Antibiotic Resistant Gene. The host cells are kept in antibiotic containing medium (antibiotic whose resistant gene is present in host along with gene of interest). Those cells which are transformed, will remain alive when treated with antibiotic whereas non-transformed cells will die of as they do not contain antibiotic resistant gene.
Blue White Screening or Insertional Inactivation
Another method to find out the transformed cells is Insertional Inactivation. This is based on the ability to produce color in the presence of a chromogenic substrate. In this, a recombinant DNA is inserted within the coding sequence of an enzyme, β-Galactosidase. Beta-Galactosidase breaks galactose into lactose. If a gene is inserted into this region, β-galactosidase will not be able to convert galactose into lactose. This results into inactivation of the enzyme, which is referred to as insertional inactivation. The presence of a chromogenic substrate gives blue colored colonies in non-transformed cells. Presence of gene of insert results into insertional inactivation of the galactosidase and the colonies do not produce any color, these are known as Recombinant Colonies.
Fig.2. Blue white screening
Watch this Video for more reference
View courses by askIITians


Design classes One-on-One in your own way with Top IITians/Medical Professionals
Click Here Know More

Complete Self Study Package designed by Industry Leading Experts
Click Here Know More

Live 1-1 coding classes to unleash the Creator in your Child
Click Here Know More